


Jamberoo (former JMolEditor) is a program for displaying, analyzing, editing, converting, and animating molecular systems.
A program is in constant development to improve the existing code and to add new functionality.
Jamberoo is triple licensed; you may distribute it under an Apache2 style license or the GPL and LGPL. You may select whichever license suits your needs.
Jamberoo's SourceForge.net Subversion repository can be checked out through SVN using following URL:
https://jbonzer.svn.sourceforge.net/svnroot/jbonzer
Jamberoo on the Sourceforge, Google group
Development of Integrated Environment for Computational Chemistry and Molecular Modeling, Presentation in Sydney University, 22/08/2007 (PowerPoint file)
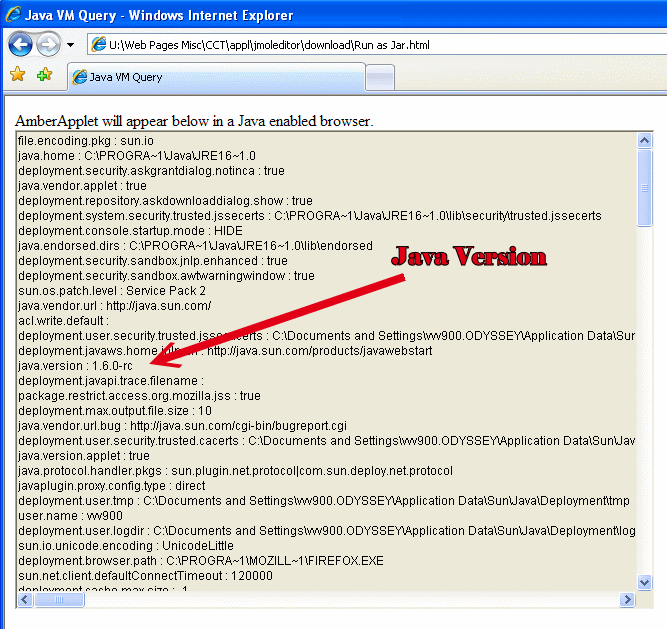
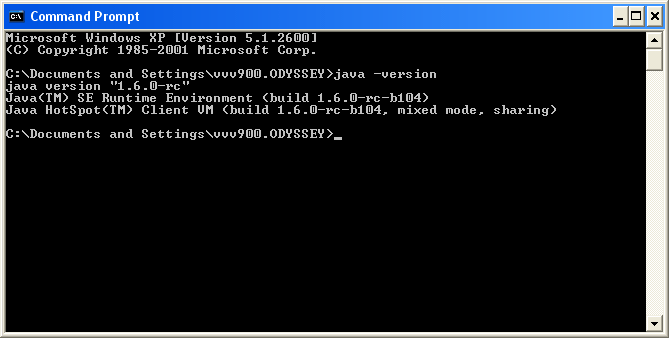
$ java -version
java version "1.4.2"gcj (GCC) 3.4.6 20060404 (Red Hat 3.4.6-3)
Copyright (C) 2006 Free Software Foundation, Inc.
The Java Web Start allows application to be started directly from the Internet using a web browser. Alternatively, application could be started from local computer using JNLP file (Java Network Launch Protocol file) which is just about 2Kb in size.
The Java Web Start caches the entire application locally on user’s computer. Thus, any subsequent launches are almost instantaneous as all the required resources are already available locally. Every time the user starts the application, the Java Web Start component checks the application's website to see if a new version is available, and if so, automatically downloads and launches it. Thus the Java Web Start guarantees that the user is always running the latest version of the software eliminating complicated installation or upgrade procedures.
Start Jamberoo using Java Web Start from a web browser: Jamberoo (with in-built help, ~11 Mb), Jamberoo-nohelp (no in-built help, ~6 Mb).
Firefox gives you two choices:
1) Selecting "Save to Disk" radiobutton you can save JNLP file to your hard drive and then to start Jamberoo by double-clicking on JNLP file.
2) Alternatively, you can start Jamberoo from browser using Java WebStart:
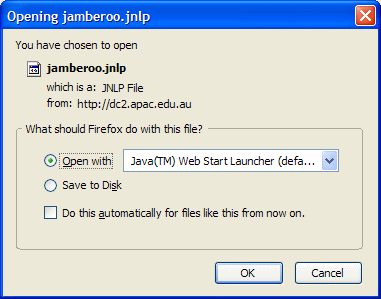
Java Web Start might show a dialog
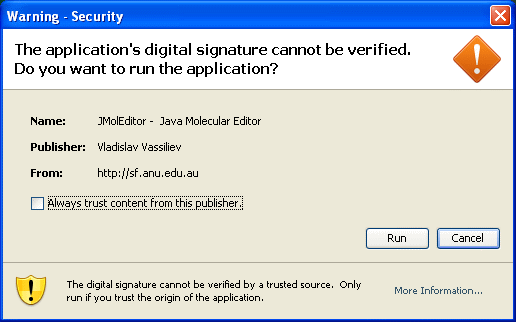
Press "Run" button to start Jamberoo (also you might want to check "Always trust content from this publisher" checkbox )
-Xmx128m option specifies the maximum size, in megabytes, of the memory allocation pool. This value must a multiple of 1024 greater than 2MB. Append the letter k or K to indicate kilobytes, or m or M to indicate megabytes. The default value is 64MB. The upper limit for this value will be approximately 4000m on Solaris 7 and Solaris 8 SPARC platforms and 2000m on Solaris 2.6 and x86 platforms, minus overhead amounts. Use -Xmx256m or -Xmx512m for studying large systems or for visualization of many molecular orbitals.
Send all questions, suggestions and comments to Vlad (vvv900@gmail.com)
Dr. Vladislav Vasilyev
National Computational Infrastructure (NCI)
The Australian National University,
Canberra, ACT, 0200, Australia
[Home] [Applications] [Applets] [API] [E-mail me]